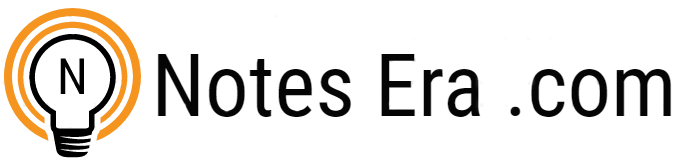Bacterial Cell Structure
Bacterial cell structure is very simple although they are very complex in behavior. They show the most extensive metabolic diversity. Electron microscope can only reveal the detailed structure of bacterial cell. It consists of following structures :
1.Glycocalyx :
- It is outermost part of cell envelope (Glycocalyx , cell wall, plasma membrane )
- Represented by either slime layer or capsule
- Slime layer is composed of dextran, dextrin and lavan sugars and protext the cell against desiccation and loss of nutrients.
- Capsule is made up of polysaccharides and D-glutamic acid, it provided gummy or sticky character and virulent property to the cell.
2.Cell Wall : it is present outside the cell membrane and is a rigid structure. Due to its rigidity, it protects the internal structures of the cell and provides shape to the cell. However, its main function is to prevent the cell from expanding and bursting because most bacteria live in hypotonic environments , and are likely to take in much water and eventually burst.
The cell walls of almost all the eubacteria (true bacteria) are made up of peptidoglycan, also called murein or mucopeptide. It is found only in prokaryotes. As the name suggests, the peptidoglycan consists of two components – a peptide portion which is composed of amino acids connected by peptide linkages, and a glycan or sugar portion. The glycan portion, which forms the backbone of peptidoglycan , is composed of alternating units of amino sugars N-acetyl- glucosamine (NAG) and N-acetylmuramic acid (NAM) joined together by β-1, 4 linkages. The peptidoglycan chains are laterally linked by short chains of four amino acid which are attached to N-acetylmuramic acid residues. The four amino acids of this tetrapeptide are D- alanine, L-alanine, D-glutamic acid and L-lysine (in Gram + ve bacteria ) or diaminopimelic acid (in Gram – ve bacteria). The tetrapeptide chains are also interlinked by a peptide braidge between the carboxyl group of an amino acid in one tetrapetide chain and amino group of an amino acid in another tetrapeptide chain. The cross linkages can occur between tetrapeptides in different chains, as well as between adjacent tetrapeptide chains. As a result , peptidoglycan forms a rigid, multilayered sheet.
Another component, teichoic acid , an acidic polymer consisting of a carbohydrate (e.g., glucose), phosphate and an alcohol is found in cell walls of Gram +ve bacteria. Teichoic acid has several functions such as binding metals, acting as receptor sites for some viruses and maintaining cells at low pH to prevent degradation of cell walls by self-produced enzymes. The walls of Gram Positive bacteria contain very little amount of lipids.
The cell walls of Gram negative bacteria are much more complex. The peptidoglycan layer is very thin making up only 10% or less of the cell wall. However, the most interesting feature is the presence of an outer membrane that covers a thin underlying layer of peptidoglycan. The outer membrane is a bilayered structure consisting chiefly of phospholipids, proteins and lipopolysaccharides (LPS). The outer membrane serves as a barrier to prevent the escape of important enzymes from the space (periplasmic space) between the cytoplasmic membrane and the outer membrane. It also prevents the entry of various chemicals that could damage the cell. It acts as main surface antigen in cell wall. However, permeability of outer membrane to nutrients is provided by proteins called porins which form channels in the membrane through which substances of hydrophilic nature and low molecular weight can diffuse.
Christian Gram (1884) developed a staining method for bacteria , using Gram sain (crystal violet). On the basis of satiability with Gram stain, bacteria are classified into two groups; gram positive and gram negative.
3. Surface Appendages : These include flagella and fimbriae (or pili)
4. Flagella are long, fine, wavy, filamentous appendages that protrude through the cell wall, responsible for the motility or bacteria. These are much thinner than the flagella or cilia of eukaryotes.